Scientific communication relies on clarity, specificity and universality. In this blog I explain how communication between medical tweeters is held back by a lack of clarity in hashtag choice, and by the absence of a “fuzzy search” feature in Twitter. I explore lessons from the way that medical research papers are categorised (MeSH headings) and propose options for improving medical tweeting, helping people to look beyond their usual social media bubble. I also demonstrate ways to visualise intentions vs reception for hashtags in two topical issues using word clouds.
I have written this as a blog, because I wanted to include a more reflective exploration of this topic than I could in a traditional medical paper. Hopefully, with the contribution of other medical tweeters, the ideas presented here can be developed into a peer reviewed paper in a medical journal.
The title and circumstances of my first proper literature review at medical school have been lost in a fog accumulated over 30 years. What I do remember, however, is the room at the front of the Erskine Medical Library on George Square, Edinburgh, where we had to book an appointment to perform a literature search. These were sought-after slots, with careful planning required in advance to make sure you used the brief time available efficiently. The stopwatch was on, and if you were quick you would be able to get your hands on the single copies of the papers you were looking for. I recall weighty tomes packed full of possible Medical Subject Headings (MeSH), a microfiche reader (I think) and a computer with that quarter’s CD of data. There was a dot matrix printer that noisily and slowly spooled out lists of authors, journals and titles – and abstracts if you were lucky – from a pile of fanfold paper with tractor holes on detachable margins. You used Boolean operators, wildcards and truncations to refine your search, expanding and contracting the outputs to produce a realistic list of papers for your project.

You then descended to the journal section of the library where decades of journals were filed meticulously. These were either bound in hardback, or loose and awaiting their eventual trip to the binders. (If you were unlucky, they were already on that trip). On the way down the stairs, I would unconsciously remove the perforated edges on the printed sheets as I started to read the outputs. I suspect everyone else did the same, but I never saw the ticker tape littering the well-worn carpet on routes to the more popular sections. A quick scan of the journal titles at the end of the rows of metal shelves allowed you to score off journals that the library did not stock – typically the most promising of titles for the esoteric topics selected. As you progressed through the years at medical school you started to know the available journals by heart, the years stocked, the change of journal titles and mergers through the decades. I would gaze wistfully at exotic titles such as Acta Scandinavica Neurologica, imagine myself exploring Ann Arbor and other previously unheard-of locations, and speculate on the nature of discussions at the Proceedings of the National Academy of Sciences. Recognising patterns in the printed output you picked your way along rows oppressive with their low ceilings and muted volumes of blue, green and purple, gauging how many you could realistically sift through before the next lecture or tutorial, and how many you should photocopy.
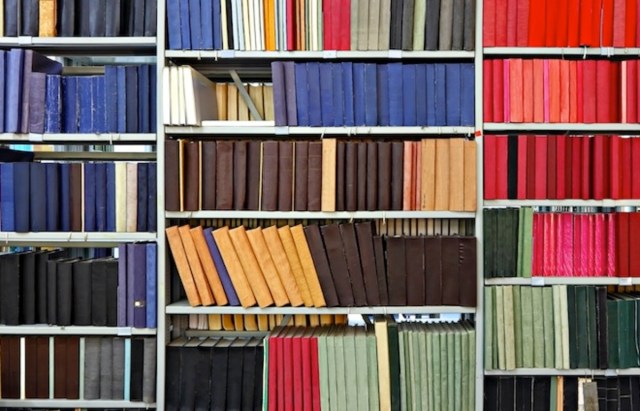

There were favourite locations to sit and study in this familiar dungeon: quiet nooks which were all too often reserved by somebody else’s pile of journals. There were also chance encounters with course mates, and discussions that had to be kept muted however welcome the interruption. In the journals there were as many blind alleys as discoveries. If you were lucky you might find a key author that had been missed in the search, or a themed issue on your specialist topic, both of which could help unlock the topic. Serendipity and curiousity brought opportunities and challenges in scoping the project, with new avenues and highly technical papers causing distractions, welcome or otherwise. Discarded volumes were placed in trolleys at the end of rows, ready for refiling by the small band of librarians, presumably because the students couldn’t be trusted to maintain order.
Judging the time available and the length of queue at the photocopiers was a fine art, frequently miscalculated. The cost of photocopying focused your mind on the relevance of the journal paper to your work, and whether the printed abstract you already had might suffice. Short deadlines, hazy objectives at the start of a project and limited library opening hours frequently meant that you photocopied more than you needed. You left the library with a bundle of papers, hoping that you had not missed a crucial page in the process, ready for long evenings and weekends of reading, thinking, analysing and writing to really begin. I know now that such work should follow a structured approach as detailed in the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) framework (introduced as QUOROM in 1996, updated in 2009).

Each of us will have a version of the search for medical knowledge at different stages of our careers. Describing the detail depending on year that you started university, and where you studied, would highlight the speed and nature of change in the retrieval and dissemination of scientific knowledge over the past 50 years. Performed properly (i.e. not just a Google search), the common factor across decades of medical literature searches is Medical Subject Headings (MeSH), introduced in the 1960s, with origins two decades earlier in the Quarterly Cumulative Index Medicus (1940). MeSH currently includes around 27,000 descriptors.
Now, of course, the whole process of searching for literature is performed on computers, mobile phones and tablets, with the full text available instantly, to unlimited users, though often you will require institutional access to read the full paper. You can choose your database depending on the topic – e.g. social sciences, laboratory, clinical. You can search in different ways, including citation searches that look for papers that cite a publication of interest. Some databases have options that allow “fuzzy searches” that look for similar terms. These in turn can help return to refine searches in databases that follow a more rigid approach. King’s College London has a useful summary of these and other advanced approaches to literature reviews.

Beyond this careful categorisation of scientific research, there are other factors that influence dissemination of research findings. Back in the early 1990s, I would sometimes find myself daydreaming of all the papers, journal issues and indeed whole volumes that had been stuck on shelves for years or decades, never accessed. (In hindsight, I realise that somebody must have dusted!) In my naivety I expected that I would soon be contributing to these stacks of sad unread papers, and wondered at the potential futility of the process. Little did I know that it would be over 15 years before I had a proper journal article published in a peer-reviewed journal. I don’t think that any of us would have dreamed at that point that authors and journals would be disseminating summaries of their tweets direct to their peers from mobile phones. We didn’t even have email at that point. Nowadays we take it for granted that authors and journals use social media to highlight their work. Such social media activity can be tracked by social network analysis, hashtag indexing tools such as the Symplur healthcare hashtag project, and altmetrics. Regardless of all these whistles and bells however the central indexing of medical research papers remains through MeSH (note: Google Scholar cannot compete with MeSH for efficiency and completeness in identifying scientific articles).
I have been reflecting on these points while I round off my social media analysis of medical topics. Over the past 4 years, my interest initially piqued by complex social network analysis maps and social media statistics shared at medical conferences, catalysed by discussion with clinical and public health colleagues across the world, and then driven by a desire to turn 1,000s of hours of analysis into peer-reviewed papers, I have authored/ co-authored 15 published papers on different aspects of the role of social media in disseminating clinical and public health messages, and am waiting on the outcome for several more articles. After the painful process of submitting some of these papers to multiple medical journals I have become reacquainted with the branching structures that inform MeSH headings, not dissimilar to the clinical coding systems familiar from epidemiological research using hospital medicine and general practice data. The process of describing a piece of work in very specific headings, to allow categorisation in a future literature search, is an essential part of disseminating medical knowledge. In a circular way I have learnt that social media platforms and users could potentially learn a lot from the careful processes used in generating and updating MeSH headings, though with considerably fewer descriptors.
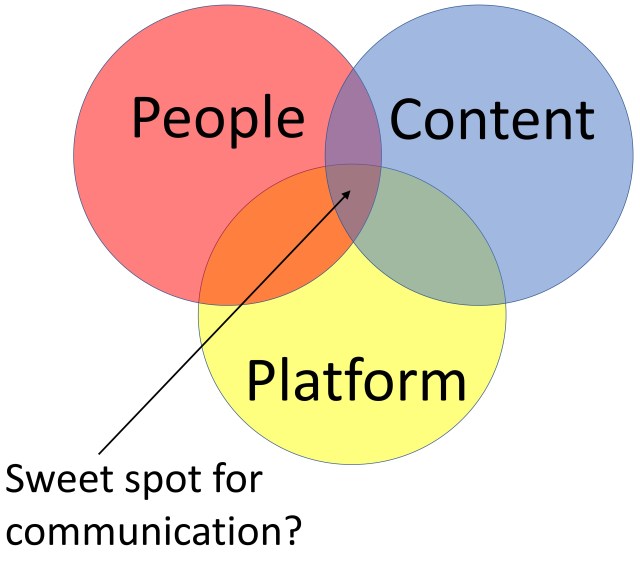
There are a number of important limitations in the use of social media for disseminating – and searching for – medical research. Superficially, social media seems straightforward: you surely just need people producing content on a social media platform for communication (Figure 1). The people may be working individually, in organisations or networks. The content can be simple text, hashtag(s), links to other information including URL to the paper and other social media users (e.g. journal, co-authors, opinion leaders in the topic), and media (e.g. a visual or video abstract). The social media platform for science and medicine will typically be Twitter. Communication results from the tweet being seen, disseminated by retweeting, replied to and quoted. There might be warm words from followers and a wider audience, discussion between co-authors, questions about methodology and findings, some debate or dispute. If the communication contributes to further understanding then it might even be classified as dialogue. The flurry of activity can be mapped and the original tweeter can see how many times the tweet has led to others clicking through to read the paper using Twitter Analytics. The URL to the journal paper can be searched to see how many times others have referenced the paper on Twitter. Prominent tweets about the research that have been quoted by others can be identified by searching for the URL of the tweet in Twitter search. It’s good to see the work being discussed – you’re left with a little serotonin hit. The experience can potentially become quite addictive.
| A hashtag, which starts with the hash symbol #, is a type of metadata tag used on social networks such as Twitter and other microblogging services. It can be a word or phrase, without spaces or punctuation (though some symbols are OK).
Examples include #Medicine and #StaySafeStayHome. |
The problems with social media come when you try to unpick this apparently simple and direct form of communication.
- First, most people are not on Twitter – it is used by around 22% of adults in the US for example, and many registered users are not all that active. However, other public platforms are not well suited to communicating science and clinical content. I also wonder how much of the peer-to-peer discussion that used to happen on Twitter is now happening in private on WhatsApp, avoiding the relentless bot activity that circles around discussions about tobacco control, vaping, vaccination and climate change.
- Second, while the content is often quite engaging, it can be difficult for the message to find its audience, and indeed for the audience to find the tweet, even though the tweeter might try to help by using hashtags. This places a major dampener on communication. Hashtags are the focus of the rest of this blog, because they are very important in the way that information is accessed on Twitter – they are commonly used to flag up key areas of interest in a tweet, and are easy ways to navigate between tweets. Compared to the clear structures and specific terms used in MeSH, hashtags are frequently sub-optimal for finding content, and this is compounded by the fact that Twitter does not have a “fuzzy search”. If you search for #CardioTwitter you will not find related terms (e.g. #Cardiology or #Cardiologist), mis-spelt words or even slightly longer variants (e.g. #CardioTwitterUK). The same is true for conference hashtags (e.g. #ACC19 vs #ACC2019), and that is even before we think about the confusion that arises when different events use the same hashtag. As new popular topics emerge the hashtags start to proliferate rapidly. The social media activity around COVID-19 is a case in point (Figure 2), with multiple hashtags used interchangeably e.g. #COVID19, #COVIDー19 (which acts as a hashtag because “ー” is not “-” which interrupts a hashtag), #CoronaVirus, #COVID, #Covid19UK, #SARSCoV2 and many others (including undesirable hashtags that promote unhelpful messages about virus origins – more on that later). Twitter does not police or curate the generation of hashtags, does not signpost you to look for other related terms, and does not flag up the gaps in your search.
- Finally, continuing on the theme of communication, while you can expand replies, follow threads, search for quoting tweets, each step takes time and technical knowledge, as illustrated in this study of a surgical conference in 2019. The original tweeters do not always know what has happened in the discussion below their tweet let alone the potential wider audience interested in a topic.
| An example of the fragmented tweeting that occurred during the early stages of the COVID-19 pandemic (Figure 2). Writing in 2014 NodeXL and Pew Research described this pattern as a combination of “community clusters” with multiple additional disconnected participants tweeting in isolation (they use the term “brand clusters”). This is a useful graphical representation of the “social media bubbles”, also known as “filter bubbles”. Such a pattern of tweeting would have a very narrow central zone in Figure 1 (the “sweet spot).
|
These issues are a challenge when looking for recent scientific content, which is often shared first on social media, sometimes a considerable time before the final paper appears in official scientific databases. This is frustrating in general reading, but is a major issue when attempting to complete a PRISMA flowchart – specifically the top right box: “Additional records identified through other sources” (see diagram above).
Presumably Twitter doesn’t mind if content is hard to find – it’s just happy that you’re tweeting, building your followers, clicking links to other tweets, going down a social media rabbit hole, feeding its algorithms, bringing in views (“impressions”) and related advertising. Twitter is a business trading on communication. However, for individual users, further steps are required in order to make it an efficient way of communicating and disseminating medical information. Unfortunately, the obvious ways to try to improve communication on Twitter do not appear to work that well. Conferences – a captive audience you would think – do not usually manage to achieve hashtag discipline, so why should we expect the same to happen in the wider world e.g. in journal clubs, clinical networks, health awareness campaigns. When typing hashtags people improvise, add things, forget things, make things up – hashtags get distorted and lose their meaning. Introducing a successful new hashtag usually takes a lot of effort and some luck, and might be short-lived even then.

Using successful hashtags has challenges too. The huge number of tweets related to the free open access meducation (#FOAMed), including radiology #FOAMrad, #MedEd and #MedTwitter hashtags are too numerous to view fully, with tens of thousand of tweets each week, so I have applied simple rules previously to list the top content. It is not immediately clear which of these hashtags to use in different circumstances – they are all used by individuals, networks and organisations, and while the balance is different by hashtag, there is a mix of medical politics, evidence-based practice generated by peers for peers, and official resources and courses tweeted in tweets including each term.
| At the height of the COVID-19 pandemic the distinction between science and politics is not always clear. As we have learnt about the SARS-CoV-2 virus we have understood the symptoms, presentation, modes of transmission, differences in susceptibility, variation in morbidity and mortality and the role of personal protective equipment and the level of lockdown. These are topics that combine epidemiology and public health, virology, healthcare procurement and politics, exposing differences between countries and healthcare settings. Unfortunately, calls for actions to protect public and healthcare workers also potentially attract the same rabid bots that disseminate disinformation about other public health topics. |
The FOAM movement has helpfully defined the intentions behind their approach: constantly evolving, collaborative, [crowd sourced] and interactive open access medical education resources, distributed on the web with one objective — to make the world a better place. This includes “medical education” in the description, which creates overlap between #FOAMed and #MedEd.
The term “Medical Education” can be viewed in a number of different ways, from process to a whole discipline. One useful explanation is that it “consists of training aimed at ensuring physicians acquire the competencies, skills and aptitudes that that allow them to practice professionally and ethically at the highest level. All physicians, the profession as a whole, medical faculties, educational institutions, and governments share the responsibility for guaranteeing that medical education meets a high quality standard throughout the medical education continuum“. While acknowledging the individual physician contribution to Medical Education, perhaps we should use the #MedEd hashtag when there is a more formally structured and top-down approach to learning.
I view #MedTwitter more as a rallying hashtag for communication between clinicians, e.g. sharing a concern, a request for help, perhaps even a joke. However, a quick review of recent tweets using this term suggests a wide range of usage.
Listing the hashtags used in recent meducation tweets is instructive, providing further evidence on the need for hashtag discipline. The following word clouds list 1,000 hashtags each: Figure 3a) the most tweeted hashtags overall; 3b) the most retweeted hashtags overall; 3c) the most tweeted hashtags after removing all the general and COVID-19 related terms; 3d) the most retweeted hashtags after removing these same terms.
Source: Extracted using NodeXL, word cloud produced using WordArt.com. See Wakelet summary for more detail and list of top tweets.
Note: These findings are based on tweets extracted 25 April to 5 May 2020. The results will vary over time. They are provided here for illustrative purposes only.
Figures 3a and 3c therefore capture the intention to share information – the number of tweets using these hashtags; Figures 1b and 1d capture the reception of that information – the act of retweeting demonstrates that the tweet has been seen and shared by another tweeter. There is quite a difference:
- Terms such as #BatFlu and #Wuhan were tweeted commonly (Figure 3a) but were much less frequently retweeted (Figure 3b). Terms that focus on origin of the pandemic are not recommended – they are potentially inflammatory, and it is reassuring that they are only used by a minority of tweeters and are not commonly disseminated by the medical Twitter community.
- Terms such as #ECG, #FOAMcc and #POCUS were retweeted much more frequently than expected (Figure 3d) compared with number of tweets posted (Figure 3c). While #orthotwitter was the most commonly tweeted topic specific hashtag, #cardiotwitter was the most retweeted.
These are helpful comparisons, but the most striking feature of each of these figures is the sheer number of terms used for the same or similar concepts. As already explained, Twitter will not flag up or group these similar terms because it does not provide a fuzzy search term.
A separate search, this time for #CardioTwitter for the period 2-11 May 2020, allows us to describe the hashtags used for a single discipline (cardiology). From these 1,125 #CardioTwitter tweets, posted by 446 people, there were 3,647 retweets. Overall, 62 (14%) tweeters received 80% of retweets, while 158 (35%) tweeters received no retweets – both figures are typically of medical tweeting, and suggests that around a third of tweeters are posting without any impact. Apart from #CardioTwitter, popular hashtags included such generic terms as #MedTwitter (n=193), #Cardiology (n=164), #MedEd (n=152) and #CardioEd (n=135). #Covid19, #covid_19 and #covid were also among the top 20 top hashtags. Excluding these popular generic hashtags and COVID-19 related terms produces the word cloud below (Figure 4). There were 893 hashtags used (after exclusions listed) of which 143 (16%) received 80% of hashtags, and 283 (32%) received no retweets, similar to the breakdown for individual tweeters. The top tweets from this extract are listed in a Wakelet summary.

The top hashtags (Figure 4) reveal interest in
- common cardiology investigations (echo, electrophysiology (epeeps), ECG – EKG was used much less commonly in these tweets suggesting perhaps a European rather than US audience)
- procedures (right heart catheterisation, Swan Ganz, radial access, TAVR)
- cardiology networks (ESCYoung, ACVC_ESC)
- conditions and presentations (heart failure, STEMI, CHD)
There is considerable overlap between these commonly used hashtags and other less commonly shared terms – e.g. #firstecho, #echo, #hf (for heart failure), #electrophysiology, #ep, #epeep, variants of #ablate. Terms such as #AFib and #AF are used in other tweets.
The difficulty of finding the often high-quality meducation content within 10,000s tweets presents medical tweeters with a dilemma – the fear of missing out versus the opportunity cost of finding that content. This is a great pity as there is plenty of novel content to be found that is not available so quickly via other routes (e.g. traditional journals). I suspect that more clinicians would use Twitter if content was easier to find, and if their own posts were more visible for others to find. Using Twitter is more than just tweeting, just as communicating is more than just talking – it is also about listening and searching out new points of interest to reflect on and potentially explore in future discussions.
We must learn to search Twitter more effectively and efficiently, just as we learnt to search medical literature when we were first starting out. Twitter allows us to refine our searches, just as medical databases provide tools to focus in on a specific period, a particular language, context or setting. When we find content of interest we can share that by retweeting, and interact by replying and following the tweeter to see their content in the future. Interacting in this way feeds the Twitter algorithms. While we know that the platform is set up in this way to generate advertiser revenue, the by-product is to increase our visibility on Twitter and, if the content strikes a chord, build followers. This in turn increases the likelihood of our tweets being shared when we want to disseminate our original research or simply learn from interacting with a wider audience.
| Some tips on Twitter search
Take the following search (click link to see the outputs or copy and paste the text into the Twitter basic search): #covid19 #kawasaki since:2020-05-01 until:2020-05-12 geocode:43.17305,-77.62479,500km It lists tweets using both #covid19 and #kawasaki hashtags, between 1 and 11 May inclusive, originating within 500kms of New York City (the geocode term). NYC was one of the first areas to report an increase in Kawasaki Disease in children during the COVID-19 pandemic. This example is provided as an illustration of the syntax of a search, demonstrating how you can restrict the parameters of a search when the original numbers are overwhelming. Of note, there have been no tweets using the terms #covid19 #kawasaki and #cardiotwitter, which means that you need to read through tweets from a range of sources rather than being able to go direct to content posted by healthcare professionals. Note that the link to the search above takes you to a view of the “latest” tweets (all tweets listed chronologically) rather than “top” tweets (a Twitter-algorithm-selected list of tweets which curiously does not always show the most popular tweets). Drawing parallels with literature reviews, if you’re interested in a topic and have the time you will typically want to see the full range of content, so make sure you select “latest”. You can add further terms, and broaden the search for international spelling and terminology e.g. #ECG OR #EKG (the “OR” needs to be capitalised); or #immunisation OR #immunization. You can try your own searches in the Twitter advanced search function or by learning the terminology yourself, which provides access to some more advanced features. Unfortunately, unlike literature searches, truncations and wildcards are not available in Twitter searches. |
MeSH requires active management, in deciding which terms to add (imagine how they’ll go about adding COVID-19 and related terms, including innovations in clinical and public health practice). Similarly, I would argue that hashtags need direct input, at all levels of Twitter usage. Crowd-sourcing is an important feature of social media activity, and is included in the FOAM definition above. While crowd-sourcing is already useful in generating content, the approach is yet to be harnessed in identifying, supporting and refining a sustainable body of hashtags that support scientific communication. Professional societies, working with their memberships and in the academies of societies that support interdisciplinary working, could
- promote hashtag clarity (an evolving A-Z of hashtags to use)
- hashtag discipline (the use of approved hashtags)
- curation of tweets (ongoing summaries of the most popular tweets that use these hashtags – this could be in the form of the Wakelet summaries referenced earlier e.g. meducation tweets).
Individual tweeters would inform and learn from such approaches, and would benefit from the wider dissemination of their tweets and the easier retrieval of content relevant to their interests. Without such input I believe that Twitter will continue to be a minority interest, limiting its role in science communication. That is not to say that it cannot be an effective platform for communication between clinicians across the world – examples such as the #SoMe4Surgery community demonstrate strong connections on social media and the real world. We need to work out ways to expand the central zone of Figure 1 (the “sweet spot” of social media communication) reliably.
Clinician programmers also have a role to play in such work – imagine an App or Twitter browser that has hashtag support built in, both for everyday clinical tweeting and conference activities.
| Application to conference tweeting: Such an App would need to allow registration of conference hashtags in advance, and would reduce the risk of hashtag drift, where tweeters quickly start to use incorrect hashtags that are much less likely to be seen by other tweeters. Databases of healthcare conferences already exist (e.g. Symplur healthcare hashtags) but their current purpose is tracking aggregate statistics (e.g. number of tweets and tweeters and rough estimates of “impressions”) rather than informing hashtag choice within tweets. |
The App would need to be reviewed and updated regularly (e.g. witness the mushrooming of COVID-19 related hashtags that led to fragmented tweeting during the first wave of the pandemic). The App could provide a three-stage approach to hashtag choice – topic, purpose and clinical hashtags (Figure 5).
I should end by saying that hashtags are not a panacea. In studies of clinical conference tweeting, hashtags are important, but so is mentioning other tweeters (e.g. a study of a European infectious diseases conference, and an American surgical conference – in press). It is important to keep the content of a tweet balanced:
- carefully chosen words that make your point unambiguously
- an image backing this up
- mention of people/ organisations referenced or who may be interested
- a small number of well-chosen hashtags
- the evidence about inclusion of a URL (e.g. to a paper, video abstract or blog) is more ambiguous, but I think that we should include such information more – it allows us to read the evidence behind the points made.
When you study medical tweeting at scale it is clear that some people think the more hashtags the merrier (sometimes 20 or more!). However, tweets with so many hashtags are difficult to read and are rarely shared. The top 100 tweets in the #CardioTwitter extract shared above (those listed in the Wakelet summary) had a median of 3 hashtags (interquartile range 2-5), though of course this extract was defined by the fact that each tweet had at least one hashtag (#CardioTwitter). These findings back up the points made in the following flowchart.

Summary and conclusions: Scientific communication relies on clarity, specificity and universality. Currently hashtag choice in medical tweets does not typically meet these criteria, though there are examples of unifying terms including #FOAMed and #CardioTwitter. However, the large number of tweets using these hashtags means that many relevant tweets are not read or disseminated. Adding further hashtags defining the purpose and specifying the clinical detail would help tweeters focus their searches. Twitter does not have “fuzzy search” function, which means that hashtag choice is crucial. I propose an approach to hashtag choice in 3 steps – topic, purpose and clinical terms – that could bring clarity to medical tweeting, but would require the type of agreement and curation provide in medical research by Medical Subject Headings (MeSH), and the buy-in of medical societies and tweeters across the world. If you agree that this would be a worthwhile endeavour, or have other suggestions, please contact me (details below) or leave a comment.
Dr Graham Mackenzie, GPST2, Edinburgh, Scotland 13 May 2020.
This blog is also available as a PDF.

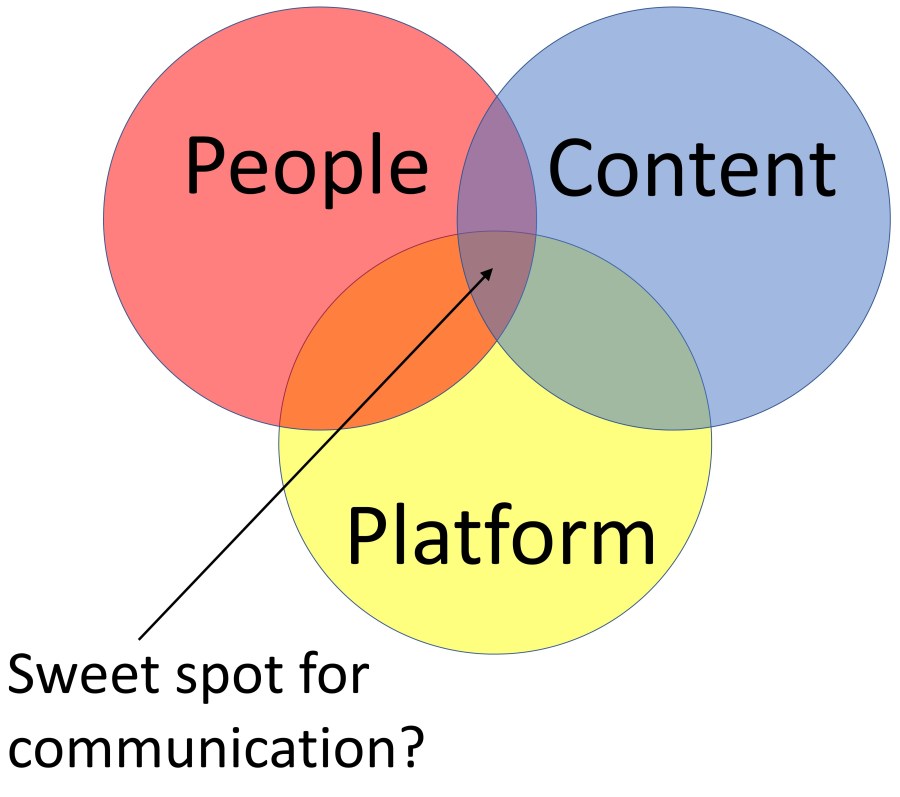





Finding the sweet spot in healthcare social media communication involves balancing informative content with engaging, empathetic interactions. It’s crucial to provide accurate, reliable health information while also addressing patients’ concerns and fostering a supportive community. Consistent, transparent communication helps build trust and credibility with the audience.
LikeLike